Microsoft CEO Satya Nadella announced on March 15 that the company's research division has published a landmark paper in Cell describing GigaTIME, a multimodal AI model that can transform routine pathology slides into detailed spatial proteomics maps of cancer cells. The breakthrough could dramatically expand access to molecular-level cancer analysis that has historically been limited by cost and complexity.
From $5 Slides to Molecular Insight
Spatial proteomics — the technique of mapping which proteins are expressed where within a tumor — is one of the most powerful tools in modern cancer research. But the imaging equipment and reagents required typically cost hundreds of dollars per sample, limiting studies to small patient cohorts.
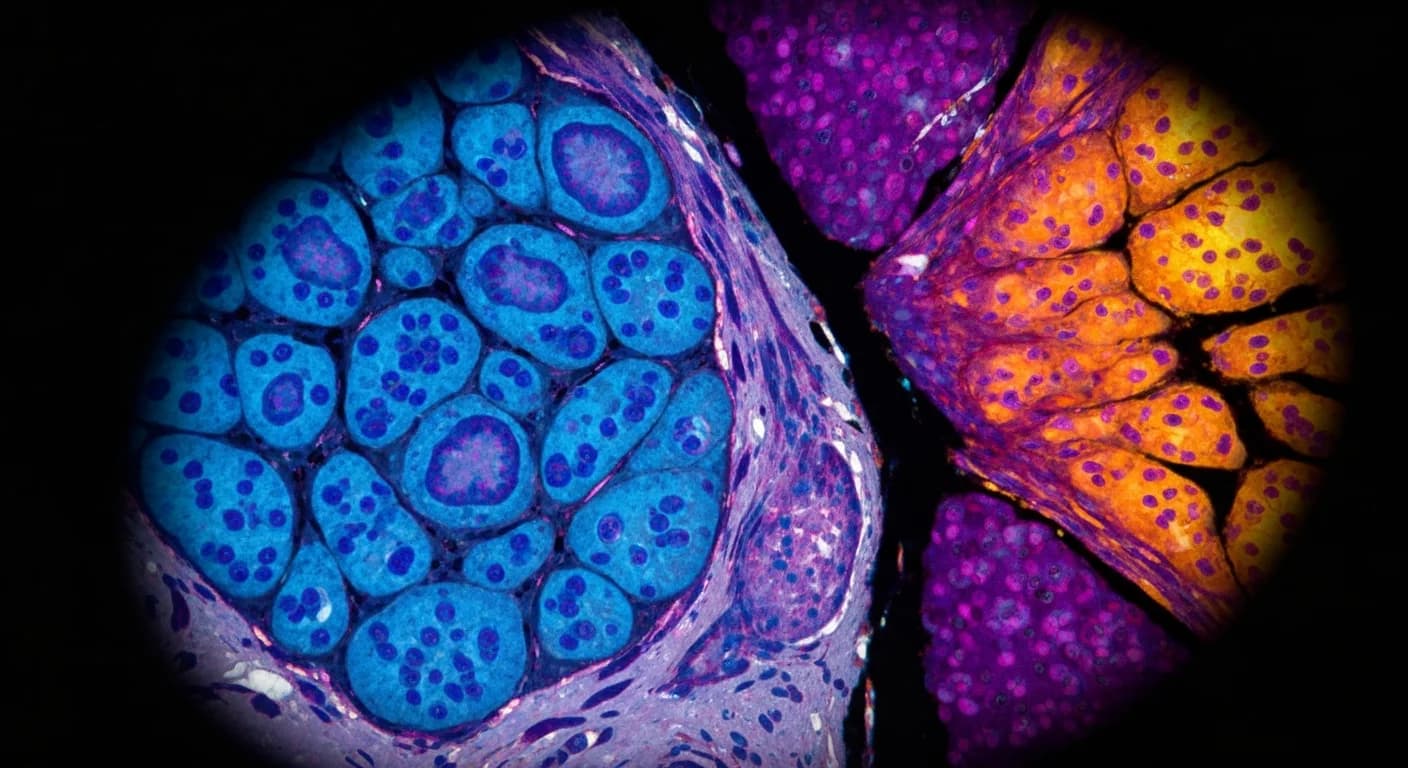
GigaTIME sidesteps that bottleneck entirely. The model takes standard Hematoxylin and Eosin (H&E) stained slides — the kind every pathology lab already produces at a cost of $5-10 per sample — and generates simulated multiplex protein expression maps. These virtual proteomics maps allow researchers to study a tumor's microenvironment at a scale previously considered inaccessible.
Scale of the Research
The numbers behind GigaTIME are striking. The model was trained on data from 40 million individual cells drawn from 14,256 patients. Across this dataset, the system identified 1,234 previously unknown protein-survival connections — relationships between specific protein expression patterns and patient outcomes that had not been documented in existing literature.
The research was conducted by Microsoft Research in collaboration with Providence, one of the largest health systems in the United States, and the University of Washington.
Population-Scale Cancer Analysis
What makes GigaTIME particularly significant is not just its accuracy on individual samples but its potential for population-scale analysis. Because it works with routine slides that already exist in pathology archives worldwide, researchers can retroactively analyze massive patient cohorts without collecting new tissue samples or running additional lab procedures.
"With GigaTIME, we can now simulate spatial proteomics from routine pathology slides, enabling population-scale analysis of tumor microenvironments across dozens of cancer types and hundreds of subtypes," Nadella wrote in his announcement.
Implications for Cancer Research
The model could accelerate drug discovery by helping researchers identify which proteins are active in specific tumor environments — information that is critical for developing targeted therapies and immunotherapies. By lowering the barrier to molecular analysis from hundreds of dollars to essentially zero marginal cost, GigaTIME opens the door for community hospitals and research institutions in lower-resource settings to contribute to and benefit from cutting-edge cancer research.
The paper is available now in Cell, with Microsoft indicating it plans to make the model available to the research community.